We reach more than 65,000 registered users in Dec!! Register Now
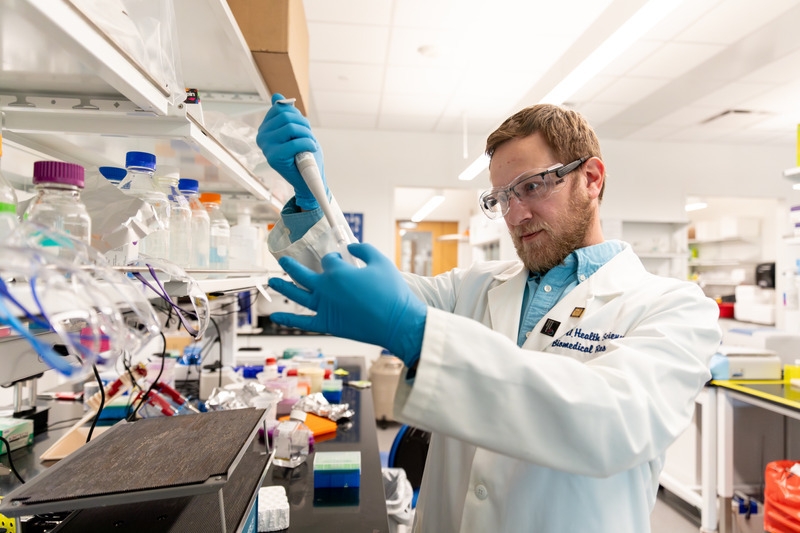

Decoding fat tissue
- February 25, 2025
- 5 Views
- 0 Likes
- 0 Comment
UD researchers find gene expression differences in fat may hold the key to targeted obesity treatment
As many as 40% of Americans are obese, putting them at an increased risk for high blood pressure, diabetes, stroke, heart disease and certain cancers, according to the CDC. New research from the University of Delaware aims to tackle the issue by investigating obesity at the gene level.
Principal investigator Ibra Fancher, assistant professor of kinesiology and applied physiology in UD’s College of Health Sciences, discovered significant differences in gene expression in adipose tissue, more commonly known as fat. Formerly considered fat storage, adipose tissue is now recognized as a vital endocrine organ. Dysfunction in the tissue is linked to significant cardiovascular and metabolic diseases.
In the study published in Physiological Genomics, Fancher and colleagues examined how diet impacts gene expression in adipose tissue using an animal model. One group consumed a diet akin to a typical high-fat, high-caloric Western diet, while the other ate a standard chow for over a year.
“We expected to see robust changes in fat, and indeed, the adipose depots in the high-fat group were much different, showing significant changes related to poor diet and obesity,” Fancher said.
Key findings
The study, funded by a federal National Institutes of Health grant to UD’s Center of Biomedical Research Excellence (COBRE) in Cardiovascular Health, found more than 300 genes were differentially expressed in subcutaneous adipose tissue (SAT), a less dangerous form of fat. In comparison, nearly 700 genes were differentially expressed in visceral adipose tissue (VAT). Visceral fat, or fat around vital organs, raises a person’s risk for significant health issues.“The comparison of VAT to SAT is stark. The expansion of visceral fat, along with its inflammatory role in obesity and metabolic diseases, is particularly severe,” Fancher said. “This study highlights the impact of obesity, which often results from a poor diet and sedentary lifestyle, on specific adipose tissues, which is very likely a major factor affecting health. That makes the affected tissue a good target for interventions to protect other systems.”
Among the thousands of genes analyzed, Fancher’s research identified four genes related to metabolism, calcium handling and inflammation that warrant further investigation.
“We’re already looking to see if these genes are worthwhile pursuits in improving adipose tissue function in obesity,” Fancher said. “They could potentially be targeted with existing drugs or spawn new treatments specifically designed to influence these genes.”
An innovative approach
Fancher worked with Bruce Kingham, director of UD’s Sequencing and Genotyping Center at the Delaware Biotechnology Institute, and Shawn Polson, director of the Bioinformatics Data Science Core at UD’s Center for Bioinformatics and Computational Biology and Delaware INBRE, as well as a research professor in the Department of Computer and Information Sciences in the College of Engineering.“Our core facilities provide access to the advanced technologies and expertise for RNA sequencing and bioinformatics that enable UD investigators to do this type of research,” Polson said. “In this project, when we analyzed the data, it very clearly pointed us to obesity-related genes and pathways that varied between VAT and SAT.”
 From left to right, Shawn Polson, director of the Bioinformatics Data Science Core at UD’s Center for Bioinformatics and Computational Biology and Delaware INBRE, and research professor in the College of Engineering’s Department of Computer and Information Sciences; Ibra Fancher, assistant professor of kinesiology and applied physiology; Mark Shaw, research associate in UD’s Sequencing and Genotyping Center at the Delaware Biotechnology Institute, collaborated on this research.Malak Alradi, a third-year doctoral student studying molecular biology and genetics, played a key role in organizing the genes into pathways to better understand their biological significance.
From left to right, Shawn Polson, director of the Bioinformatics Data Science Core at UD’s Center for Bioinformatics and Computational Biology and Delaware INBRE, and research professor in the College of Engineering’s Department of Computer and Information Sciences; Ibra Fancher, assistant professor of kinesiology and applied physiology; Mark Shaw, research associate in UD’s Sequencing and Genotyping Center at the Delaware Biotechnology Institute, collaborated on this research.Malak Alradi, a third-year doctoral student studying molecular biology and genetics, played a key role in organizing the genes into pathways to better understand their biological significance.“Before I started this research, I thought fat was the same in the body, but when I saw the RNA sequencing and studied the different genes and pathways, I realized that VAT is affected by obesity far more than SAT,” Alradi said. “Our approach shows how interconnected these processes are and why targeting specific pathways could make a difference in obesity treatment.”
Stringent statistical methods also confirmed key findings about adipose depots, including changes in metabolism and inflammation.
“That makes us feel really good about the genes we identified,” Fancher said. “It underscores the novelty of our findings.”
List of Referenes
- Malak Alradi, Hassan Askari, Mark Shaw, Jaysheel D. Bhavsar, Brewster F. Kingham, Shawn W. Polson, Ibra S. Fancher. A long-term high-fat diet induces differential gene expression changes in spatially distinct adipose tissue of male mice. Physiological Genomics, 2024; 56 (12): 819 DOI: 10.1152/physiolgenomics.00080.2024
Cite This Article as
No tags found for this post